These days, very little science occurs without someone typing at least a few lines of code into a computer. Researchers employ a variety of programming languages — such as R, Python and Bash — and software to organize their data, perform analyses, build models, and visualize results. College of the Environment scientists are no different, and that has implications for science, communication and how students will gain new computational skills in the future.
While we often associate coding with cutting-edge analyses, some researchers need programming and software to simply see and manage their data. Steven Roberts, a professor in aquatic and fishery sciences, uses DNA sequences to understand how marine invertebrates respond to environmental change by altering their physiological processes.
“My lab and I approach physiology from a genomic molecular approach — meaning sequencing DNA, mRNA and proteins to see how they vary between individuals to better inform our understanding of how marine invertebrates live. It feels like it’s mainly just moving files around and reformatting them, but we use a computer to do that.”
Roberts is also a proponent of open science — a movement to make science more transparent and accessible through shared data and methods, thus allowing for increased collaboration. In his experience, this approach to research has naturally broadened his lab’s network by encouraging prospective students and new collaborators to reach out and work together on new projects. As he puts it, “open science breaks the ice in communications in a way that pays off in the long run.”
Beth Gardner, a professor in environmental and forest sciences, uses code to model natural processes in wildlife populations based on field observations. “If we just have data it’s hard to tease apart what we observe, which has biases like detection error, from that true process — what’s really happening. We need this step of modeling to separate what we see from what’s occurring in reality,” she explains.
Method-sharing has allowed others to adopt and adapt some of Gardner’s spatial capture-recapture models — tools that estimate the number of individuals in a population from detections and redetections across space — for their own studies.
“I started developing these spatial capture-recapture models during my postdoc and those models have just blown up,” said Gardner. “They’re the base of what people use now if they’re going to do a capture-recapture analysis in terrestrial science. It’s fun to see how many people have picked them up and developed them in different directions; we keep using that framework because it’s so flexible to be able to answer all kinds of new questions.”
Code is also proving to be a key tool in science communication. Dargan Frierson, a professor in atmospheric sciences who builds simple climate models, employs code and video game software both to visualize his own research and empower others to engage a broader audience through his work with Earth Games.
“It’s really fun to make up imaginary planets and not know what to expect and see those first animations come out and weather systems just appear. The main idea behind Earth Games is to share that experience with people — the thrill of new discovery and watching this world you’ve created come alive.”
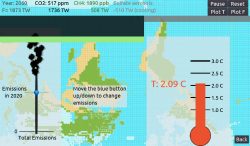
Recently, Frierson has started exploring simple web-based video game engines as a means to interactively communicate research findings. “I’ve been messing around with a video game engine called Godot that lets you make shareable, interactive visualizations with ease. I started to make these interactive game-like visualizations to aid in research communication, because instead of viewers interacting with the code they use a little slider to change parameters and watch the data model in different ways.”
Many researchers see a future where these skills are a necessity for scientists. “Ultimately if you want to do science you’re probably going to need to do some coding,” Gardner emphasizes. Her advice for students looking to learn programming languages like R is to take advantage of classes offered through the College of the Environment, like those in the Quantitative Science program.
“I always tell people that these are like spoken languages. You have to be able to have practice all the time and keep up with it so that you’re able to maintain it. It’s hard to learn any language without some application, so when you’re learning it helps to have your own dataset or question to work on.” Gardner notes, however, even seasoned researchers still frequent Google to surmount their programming roadblocks.
Roberts, who teaches an introductory data science class in fisheries, acknowledges that learning programming can initially be quite daunting for students. “I start easy with data science and what it is. Programming requires a change of mindset when you get into analyses initially. It’s easy to take for granted when you’ve been doing it for so long, but for students first learning these concepts the basics such as file and directory structure can prove to be challenging,” he explains.
Additional resources for both undergraduate and graduate students to explore include data science workshops through The Carpentries (occasionally offered on campus through the UW eScience Institute), the eScience Institute’s Winter Incubator program which pairs researchers from any department with data scientists to address a research question and the UW’s Research Computing Club, which provides access and training for the UW’s supercomputing cluster and cloud computing resources.

